Our work on the structural determination of malate synthase G (MSG) using cryo-EM and X-ray crystallography was recently accepted and published in the Journal of Structural Biology.
MSG is an E.coli protein containing 723 residues with a molecular weight of 82 kDa. We used cryo-EM to determine the structure of this sub-100 kDa MSG protein at 2.89 Å. The work was initially collected in January 2019 when I started my independent position at the Institute of Biological Chemistry, Academia. I processed the data and got a 3.4Å map in early 2019 (cryoSPARC 2.1 and RELION 3). The data was good, but I was unsure what to do as we are not the first one showing sub-100kDa cryoEM protein structure. The resolution wasn’t the best record in the world at that time. However, this is still a significant milestone as the cryo-EM facility in Academia Sinica, Taiwan, was established in mid-2018. I am a newcomer, and the data demonstrated that my group could do cryoEM data collection and analysis as perfectly as other outstanding teams.
We used cryoSPARC 3.2 to reprocess the raw data revealing a 2.89Å map. The structure was then determined. The image below presents how small MSG is compared to a 2.2 MDa ribosome structure (also determined by cryo-EM). The MSG cryo-EM structure is available in RCSB PDB, PDB ID=7YQM. I also shared the motion-corrected movies deposited in EMPIAR (ID: 11167, released soon).

We also analyzed the cryo-EM map of MSG, revealing the dynamic pattern of MSG in cryo-EM is highly consistent with what was observed using solution nuclear magnetic resonance (NMR). As the previous MSG crystal structures were ligand-bound forms, we decided to obtain a crystal structure of apo-form MSG to compare the structural dynamics of MSG in the crystal form. Our X-ray crystal structure was determined at 1.6 Å (PDB ID 7YQN) using the best synchrotron beamline TPS 05A in the National Synchrotron Radiation Research Center. The apo-form X-ray crystal structure is nearly identical to the cryo-EM MSG structure, and the b-factors in our MSG structures (two chains) are consistent with solution NMR and cryo-EM data.
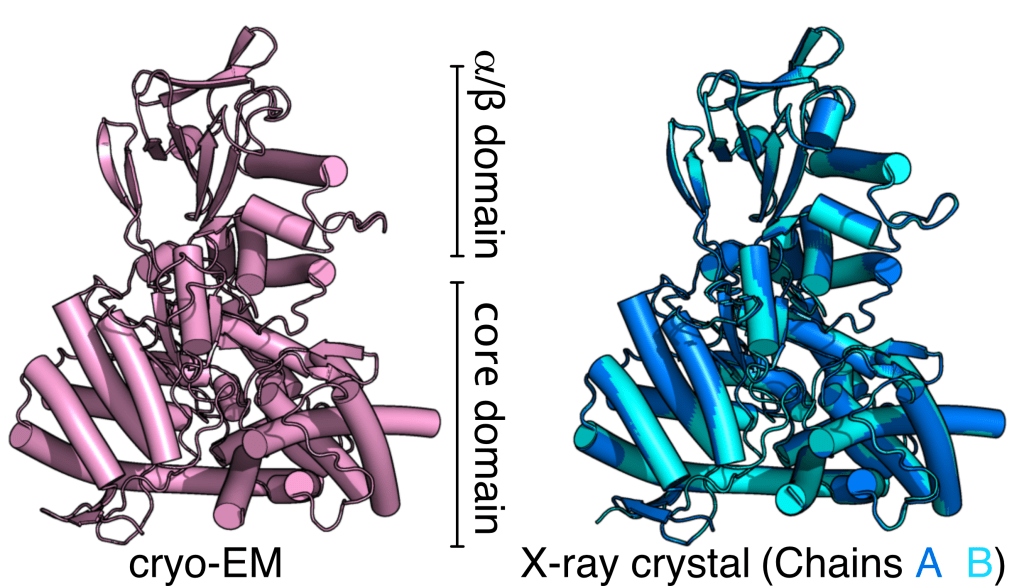
Overall, we are happy to present that cryo-EM is a powerful tool for structural determination for mega-dalton and 60-100 kDa proteins. The dynamic pattern of the cryoEM MSG map revealed by cryoSPARC’s 3D variation analysis (3DVA) fairly reflects the solution (or solution-like) dynamics of MSG observed by NMR. For more details, please go to read our manuscript in JSB.