This blog post aims to show you the resources that you can pick up one to add hydrogens for your desired structures properly. I will cover PyMOL, UCSF Chimera and Reduce in this post.
Most of time, when we download PDB files from RCSB protein data bank, the structures were determined by X-ray crystallography meaning the structures are likely hydrogen excluded. I think many cryo-EM structures are in a similar format as people generate templates from crystallographic ones. I do see more and more software in cryo-EM adding hydrogens by default.
This is very different from NMR-based structures as NMR structures are mostly calculated using NOE-based H-H distance constrains. When I want to add hydrogens to to existing crystallographic structures, there are few software available. PyMOL is a quick option but the nomenclatures of added hydrogens don’t follow IUPAC style (I will show example later). If I need to do more thorough works, for example checking consistencies between NMR and crystal structures, I prefer using UCSF Chimera or Reduce to generate proper hydrogen-added structures.
In PyMOL, the command to add hydrogen is “h_add” or users can simply click “A -> hydrogens” to add all hydrogens or selectively add polar hydrogens (same as remove hydrogens).
PyMOL>h_add ?
Usage: h_add [ selection [, quiet [, state [, legacy ]]]]
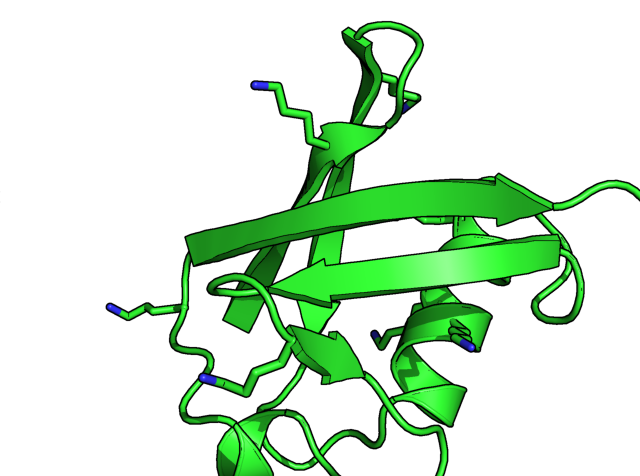
I am using 1UBQ (ubiquitin structure determined by X-ray crystallography) as the example in this post. The left panel below selectively displays lysine sidechains with sticks. Then I typed
h_add 1ubq
to add all hydrogens on 1ubq, the hydrogens are automatically displayed in the right panel below.
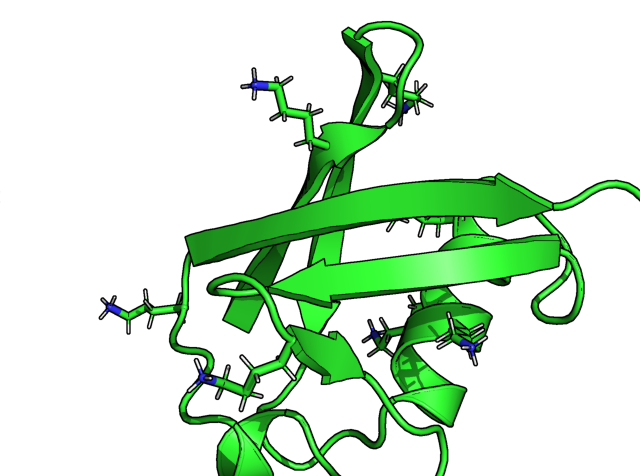
The issue at here is that the hydrogens are named H01, H02, H03 .. etc which are atypical. The standard nomenclatures of hydrogens are associated with the positions/connected atoms. For example HA (H-alpha) means hydrogen connected to C-alpha. The example below shows Lys6 sidechain NH3 are named as H13 (and H11, H12, not clicked). The right image below further shows all hydrogens (except HA) added for lysine 6 that they are numbered from HA to HZ.
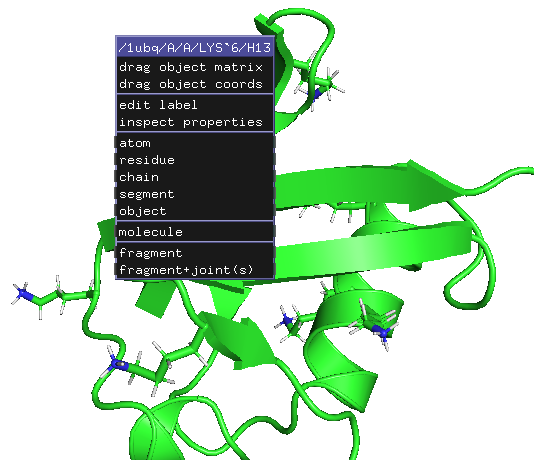
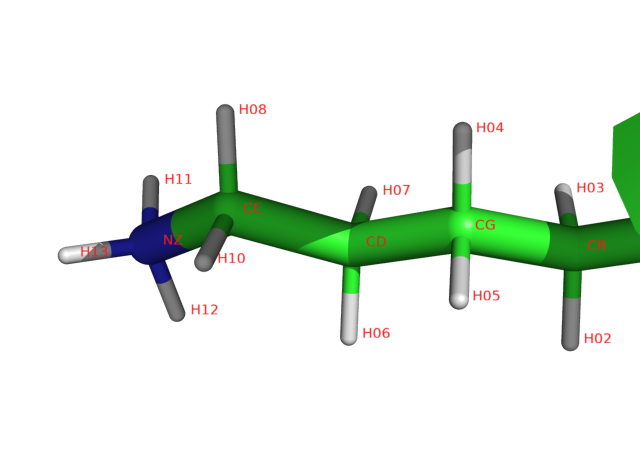
On the contrast, I often use UCSF Chimera to add hydrogens as it adds hydrogens followed by the common nomenclature. Here are the steps in screenshots for adding hydrogens in UCSF Chimera.
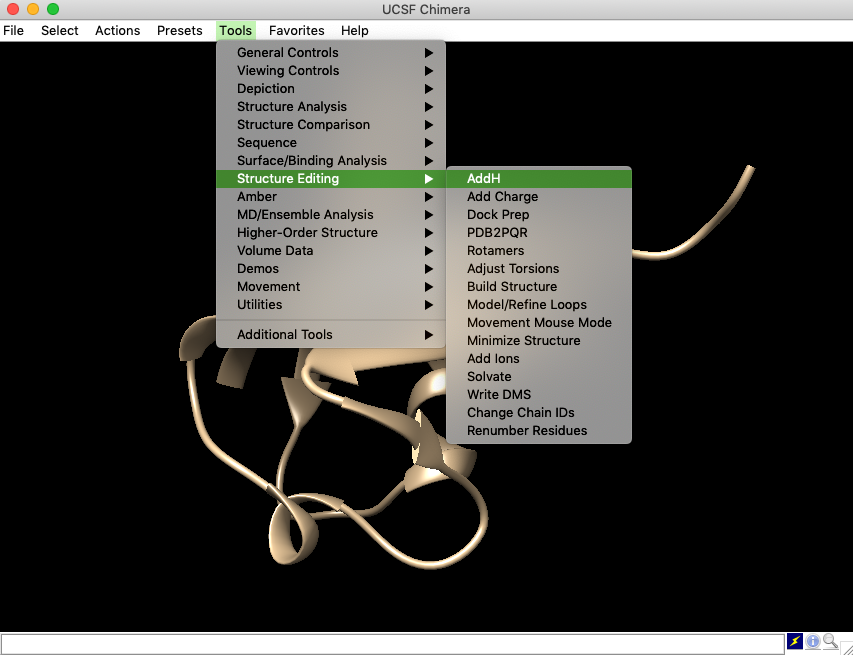
- First, load target protein structure (here is 1UBQ), then go to the tool bar, find “Tools”, select “Structure Editing” and the popped sub-panel has “AddH”. Chose it.
- After clicking “AddH”, the “Add hydrogens” window shows. If you have no particular goals, simply click “OK” button and UCSF Chimera will generate hydrogens for you.
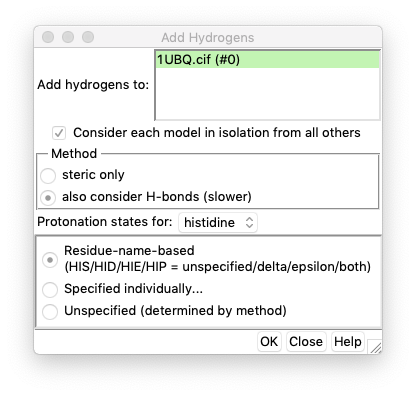
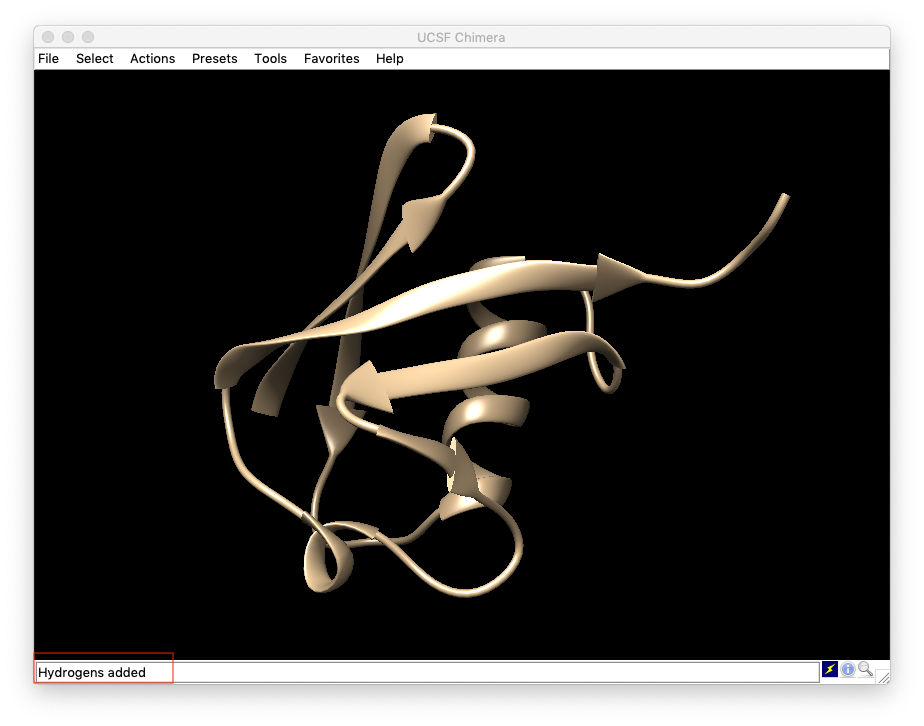
- Once hydrogens are added, the window closes. Users can check the bottom left corner in UCSF Chimera window to confirm hydrogens added (see the red boxed area in the second-to-last figure).
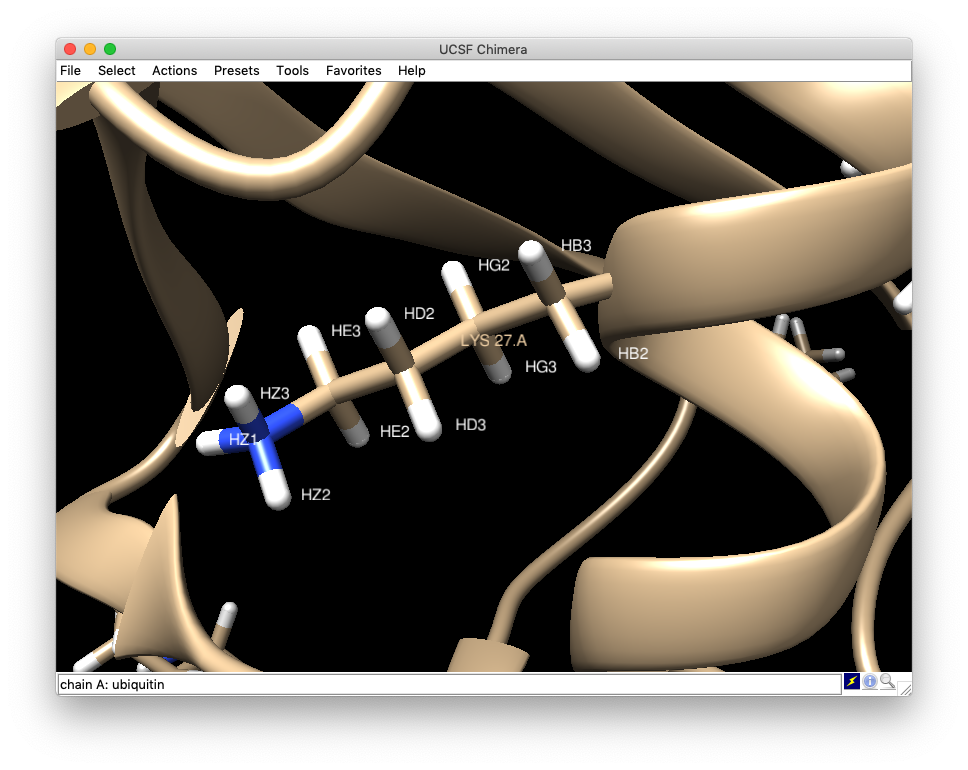
- The bottom figure shows labels of each atoms of Lysine 27 that the hydrogens are named by the positions of carbon/nitrogens from alpha, beta, gamma and so on..
- To use this hydrogen added PDB for other purposes, go to “File” and save it. Then the hydrogen added PDB file is ready for further usages.