In PyMOL, users can easily add/create or delete chemical bonds. For example, a disulfide bond is expected to be shown in PyMOL but not automatically displayed. It occurred to me when dealing with newly determined structures, especially the NMR structural cases. On the other way, we sometimes download a PDB-formatted structures and found some bonds are not appropriately displayed. Both examples can be easily handled by PyMOL using “bond” or “unbond” to create or delete bonds, respectively.
The command syntax includes:
PyMOL>bond ?
Usage: bond [ atom1 [, atom2 [, order [, edit [, quiet ]]]]]
PyMOL>unbond ?
Usage: unbond [ atom1 [, atom2 [, quiet ]]]
For deleting a bond, here I use SARS-CoV-2 main protease structure (PDB: 7BUY) as an example, which has a covalent bonded inhibitor named carmofur. Left image below is the automatic display in PyMOL (I am using 2.3) where the C07 carbon and the C145 SG atom are bonded. I used the command below to remove the formed bond. The result is shown on the right panel.
PyMOL>unbond /7BUY/B/A/JRY`401/C07, /7BUY/A/A/CYS`145/SG
Unbond: 1 bonds removed from model "7BUY".
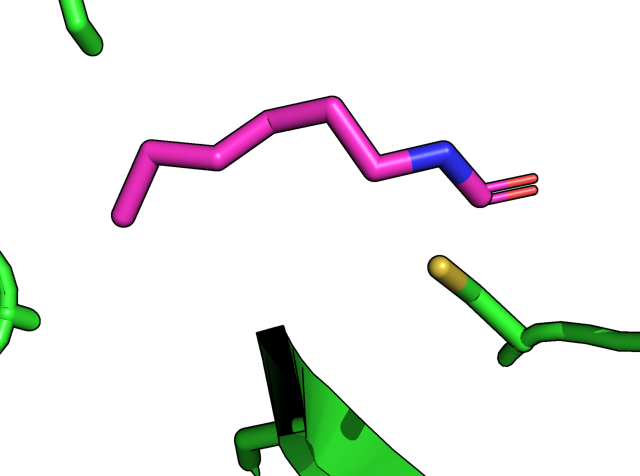


Similarly, the bond can be added back by the “bond” command.
The “bond” and “unbond” commands are not handled based on the scientific knowledge but arbitrarily manipulated by users. Users must be very clear with the reason adding or removing the bonds in PyMOL. I am showing a manipulated case below that I added a N-S bond (very unlikely happened in the protein world) in the same main protease structure.
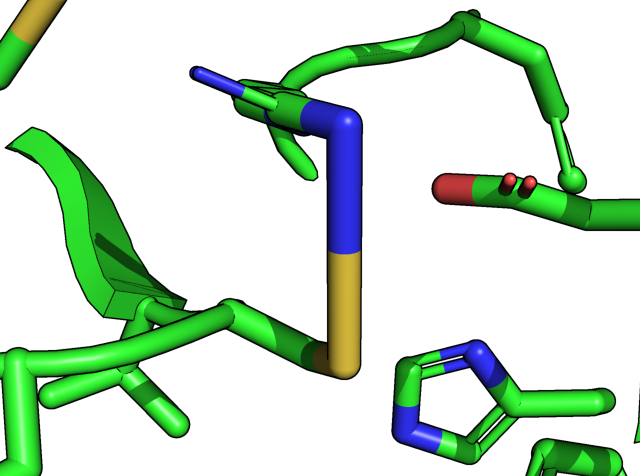
The left panel below indicates that the guanidino group of R40 (the NH2 sidechain) is 3.7 Å away from the sulfur thiol group of C85. I forced the two atom (N of R40, S of C85) forming a bond by the command below. The right panel below shows an ugly (and extremely long) N-S bond.
PyMOL>bond /7BUY/A/A/ARG`40/NH2 , /7BUY/A/A/CYS`85/SG
Bond: 1 bonds added to model "7BUY".

In short, the usage of bond/unbond must be very careful. The two commands might not be used in most of cases while using PyMOL but they are useful commands to display some bondings, especially a new ligands or any modifications on the sidechain nitrogens or sulfurs.