To analyze your interesting/study proteins, knowing distances for some key contacts are important. PyMOL provide commands to measure distance between two atoms or user can measure/list atoms at a fixed distance.
I often use “distance” (same as “dist”) to analyze my study cases. The argument of “distance” is shown below. Typing “distance ?” can show the arguments of specific commands. More details can be found at PyMOLwiki
Usage: distance [ name [, selection1 [, selection2 [, cutoff [, mode [, zoom [, width [, length [, gap [, label [, quiet [, reset [, state [, state1 [, state2 ]]]]]]]]]]]]]]]
Here I use ubiquitin (PDB: 1UBQ) to show several examples of “distance”.
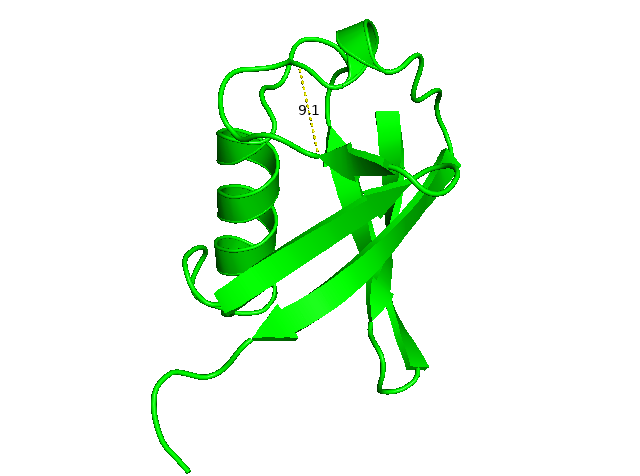
To measure the distance between the loop of 48-55 of ubiquitin. I chose residues 50 and 55 and their carbon alpha (CA). We can type command below to get the measured distance: 9.1 Å. PyMOL will show a yellow dashed line (like hydrogen bonds) and a label number to show the distance and connection.
distance d1, ////50/CA, ////55/CA, (d1 is the assigned selection object.)

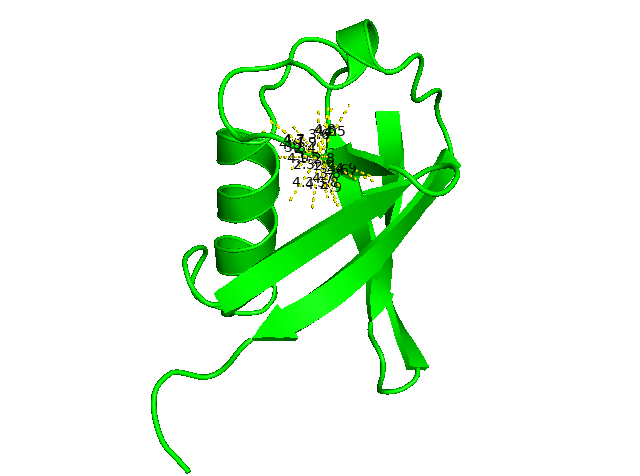
Sometimes, we want to know a list of one-to-others at particular fixed distances. For example, I want to know which atoms are up to 5.0 Å away from CA of residue 50. Use the same distance command and specific upper limit of distance “5” (cutoff in Angstrom) to display all distances matched to the selection. The left figure below is then zoomed in to the right panel to show detailed 50/CA to other atoms within 5Å
distance d5A, ////50/CA, all, 5

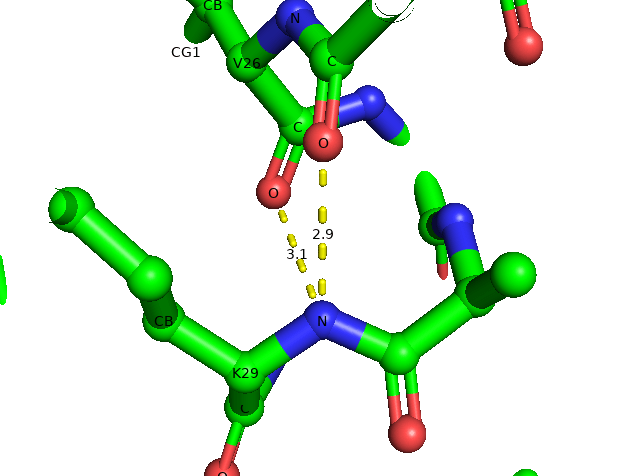
By taking the concept of distance range, we can select distances fitted to hydrogen bondings. For example, I wan to know how many hydrogen bonds formed by Lys29 backbone amide. I should specify selection for polar atoms only (mode=2). The distance command returned me two H-bonds which are K29N to N25 CO and V26CO at 2.9 and 3.1 Å, respectively.
distance hbond-29, ////29/N, all, 3.2, mode=2
Combined selection and distance is highly suggested. Users can play around the commands and think of what are the needs for the research project/data.