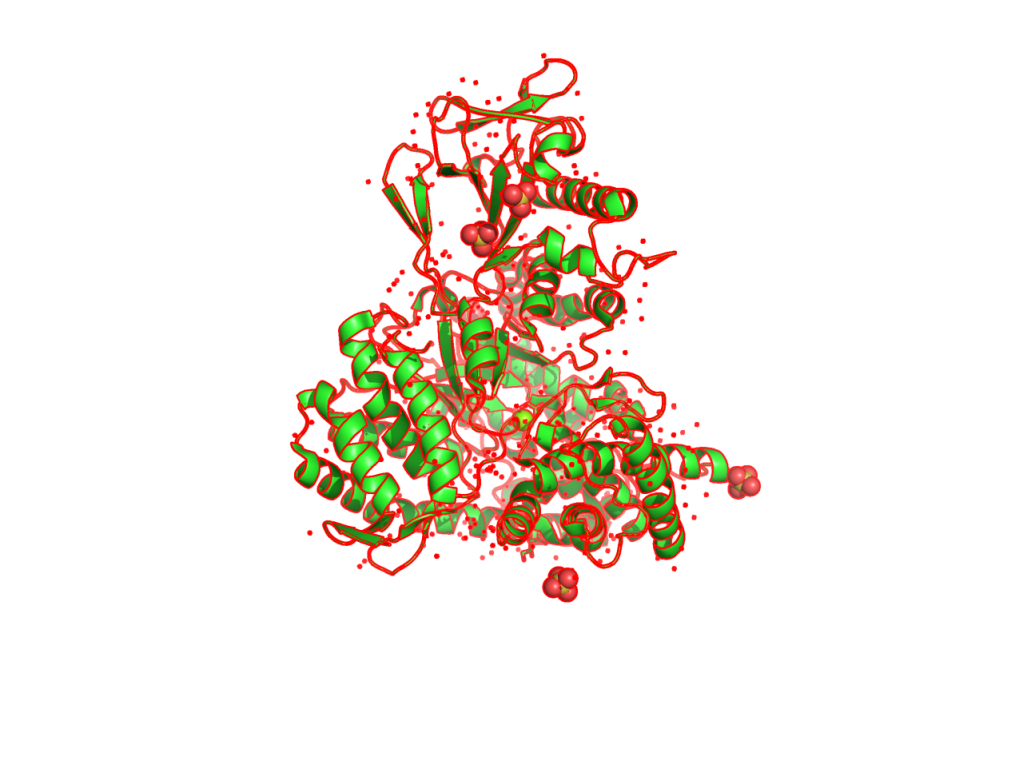
In PyMOL, one can quickly export views of molecules from “File ->Export Image as -> PNG”, however it doesn’t render shadows and other settings. To generate a nice view of protein or biomolecules, PyMOL provides “ray” command to produce more features of painting as well as size of output image.
Here, I am showing a command called “ray_trace_mode” with 4 modes to present a taste of rendered views.
(1) set ray_trace_mode, 0 –> default view of PyMOL.
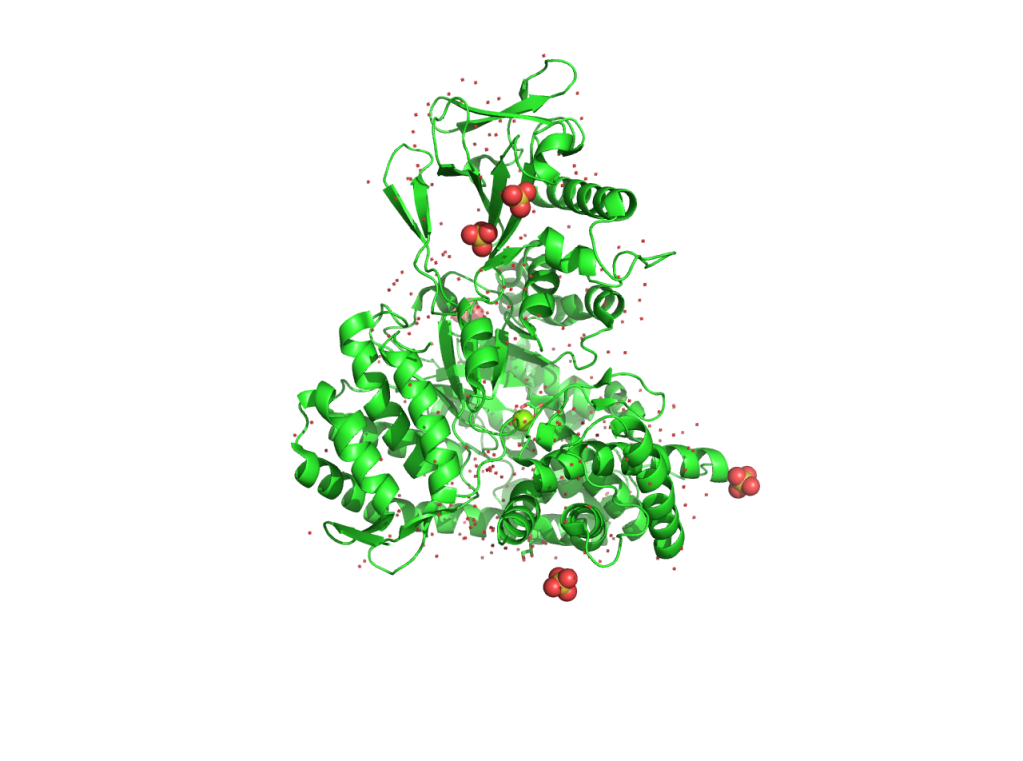
(2) set ray_trace_mode, 1 –> normal color + black outline
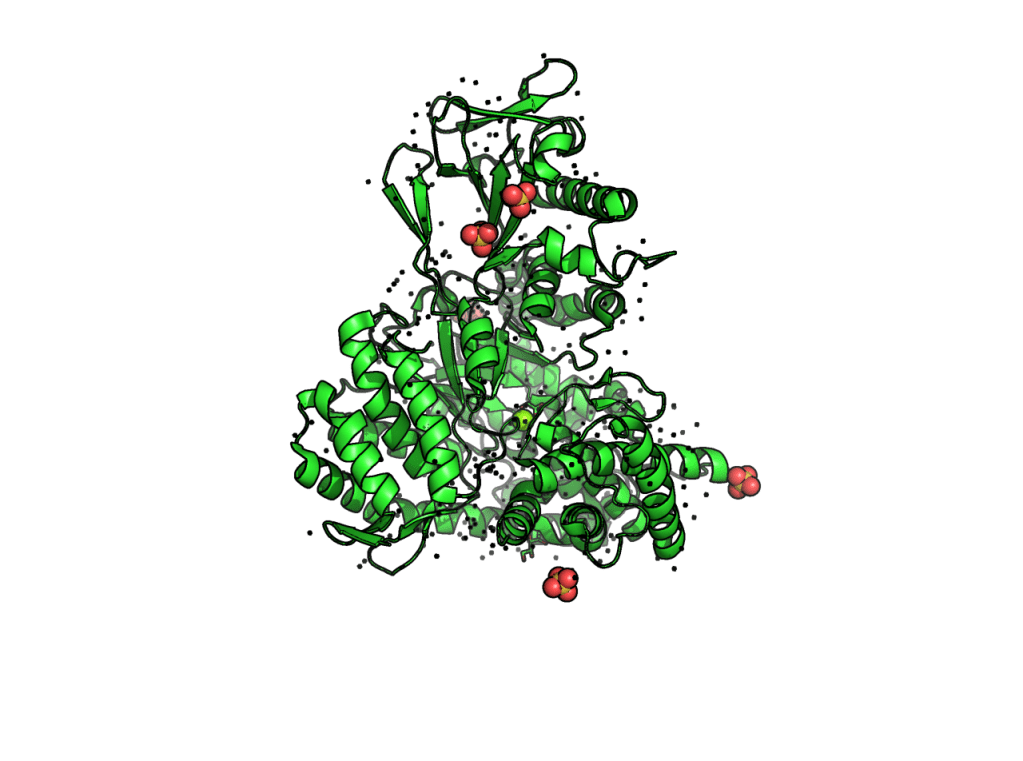
(3) set ray_trace_mode, 2 –> black outline only

(4) set ray_trace_mode, 3 –> quantized color + black outline
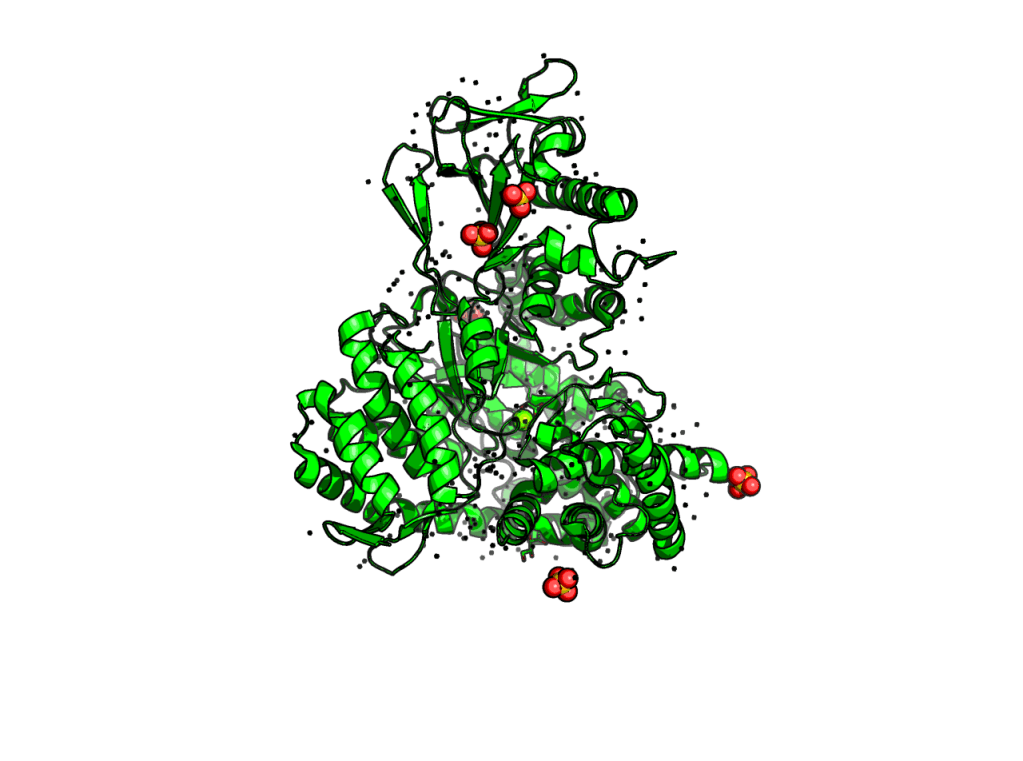
Moreover, color of “trace” outline can be defined manually.
(5) set ray_trace_color, red –> outline is colored in red