A quick show of several ways to present DNA molecules using PyMOL
Demo: 2WCC (RCSB link)
step 1: fecth 2WCC
in PyMOL 2.3.2, it automatically display cartoon view of 2WCC as:
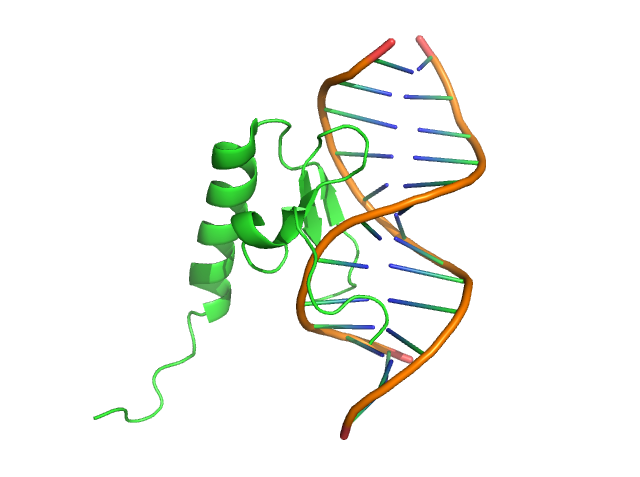
Then, we can change the DNA double strand “backbone” from “tube” to “oval” shape:
step2: cartoon oval, ///1+2
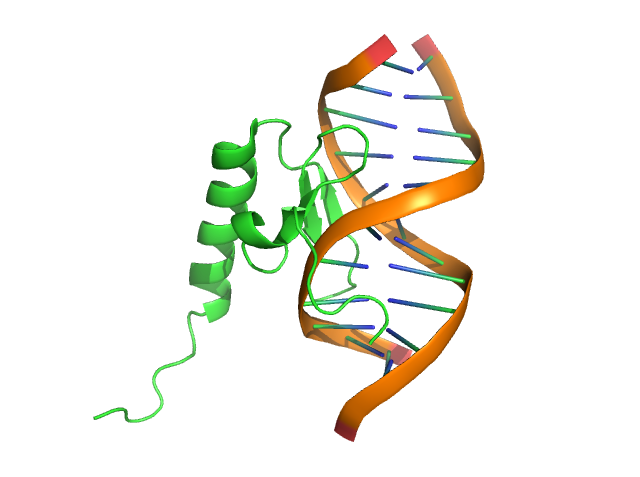
Now, let’s show the “rings” of nucleic acid and color the DNA backbone to yellow.
Step 3: set cartoon_ring_mode, 3
Step 4: set cartoon_nucleic_acid_color, yellow
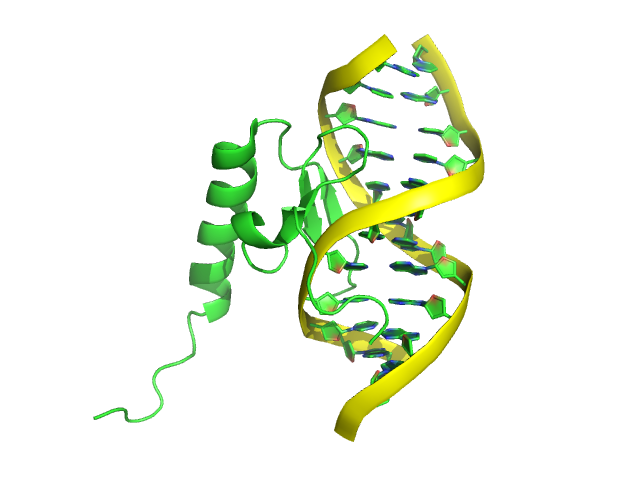
We can also further color the “ring” of nucleic acid.
step 5: color red, resn DA+DC+DG+DT
(DA, DC, DG, DT are the residue names of the 4 deoxyribonucletide)
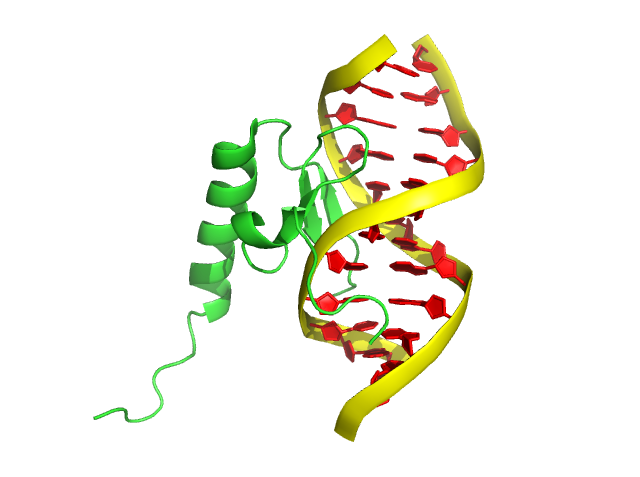
If different colors for DNA nucleotides are needs, PyMOL can easily color each pair (or individual) separately.
step 6: color blue, resn DA+DT
(in the case below, AT and GC pairs are shown in blue and red, respectively)
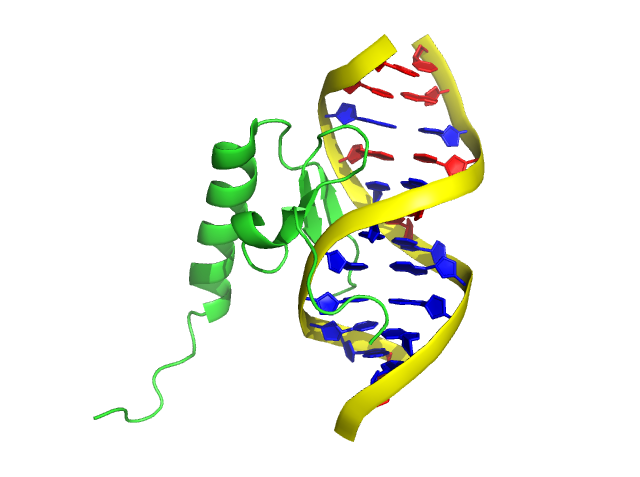
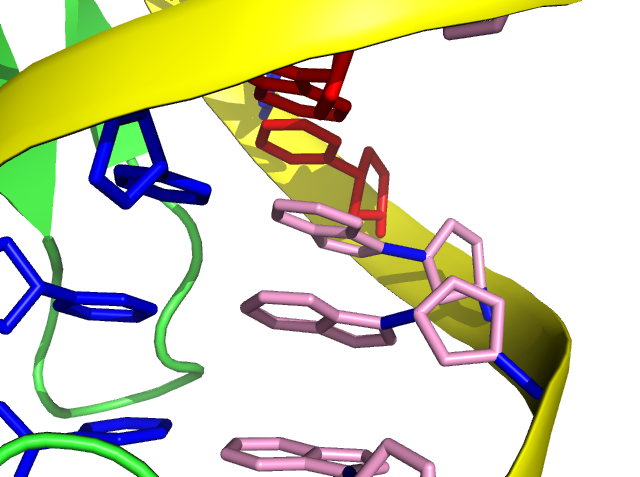
Furthermore, the colors of nucleic “rings” can be painted alone.
Step 7: set cartoon_ring_color, pink, resn DA
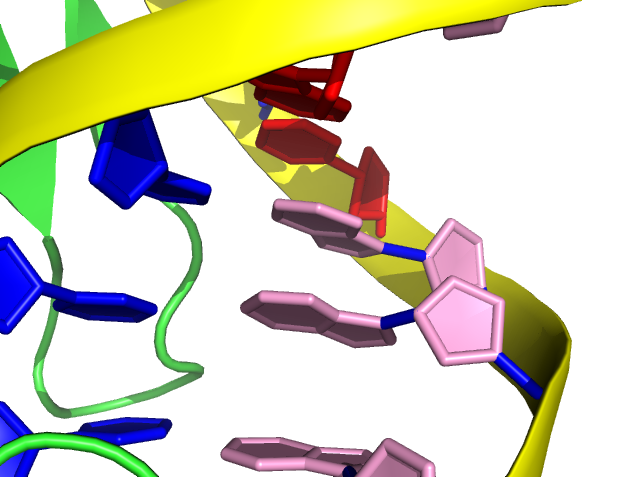
In PyMOL, filled ring is presented in default and we can turn on the transparence (0 to 1) manually.
Step 8: set cartoon_ring_transparency, 1