Background:
I did use TOPAZ back to early 2020 to get particles for single particle reconstruction. TOPAZ did good job but it took too long to get all micrographs denoised, particle train set and picking. In the end, I used blob picking in cryoSPARC to get most particles and polished junks by 2D classifications in few hours. So at that time, I was not enthusiastic to adapt AI for cryoEM data particle picking. I read a few papers recently and some of them used crYOLO‘s deep learning trained auto-picking solution to get particle and/or filament lists. Like TOPAZ, the installation of crYOLO using “conda” is very convenient. Thanks to the author’s efforts.
I actually had a few bad experiences using RELION’s auto helix picking to get fibril segments. It ended up a waste of 3-5 days to remove junks from the segment pools. The final map was much worse than manually picked fibrils. So I always spent time on fibril picking to ensure the following steps are smooth and trouble-free.
With a short trip over the weekend, I decided to play crYOLO with my fibril dataset during train transportation. The crYOLO team has done excellent tutorials; the best place to learn crYOLO. I used the box manager to pick ~50 fibrils in 10 micrographs. Trained the dataset with 300 micrographs. Training took 30-40 minutes by 4 GPU cards (Nvidia 2080 Ti, 12GB each) in my 3 random trials. Then ran crYOLO’s predict with trained model for 6000 micrographs. crYOLO returned me ~100,000 fibrils no matter definitions of mask diameter and fibril size were checked or not. The auto-picking process using 4 GPU cards (again, 2080 Ti) was about 1-2 hours. Fantastic efficiency. Most jobs were done when sitting in a high-speed train connecting back my workstation with weak iPhone 4G signals.
Now the problem is how to import the picked fibrils into RELION for further data process. I used a similar but different way described in crYOLO’s import tutorial. Thus I could check the quality of particle/filament picking.
Here is my solution:
1. Go to "STAR-START-END" folder, find out the RELION-format star files recording XY positions of fibrils. 2. run this command below rename "s/fractions/fractions_manualpick" *.star It will extend the file names with "_manualpick" for all files in this folder. 3. Assuming user has done RELION motion corrections, CTF estimation and one manual pick job was done. Copy the whole renamed files to Manual/job00x/Micrographs/ Note: assuming the image folder named Micrographs. 4. In the manual picking job, I click "Continue" button to view the picked fibrils. It works well for RELION 3.1 and newer (at least, good for my workstations). 5. This should work for crYOLO picked single particles (haven't tested yet). 6. After quick manual inspections, I was happy and ran particle extractions and 2D classification.
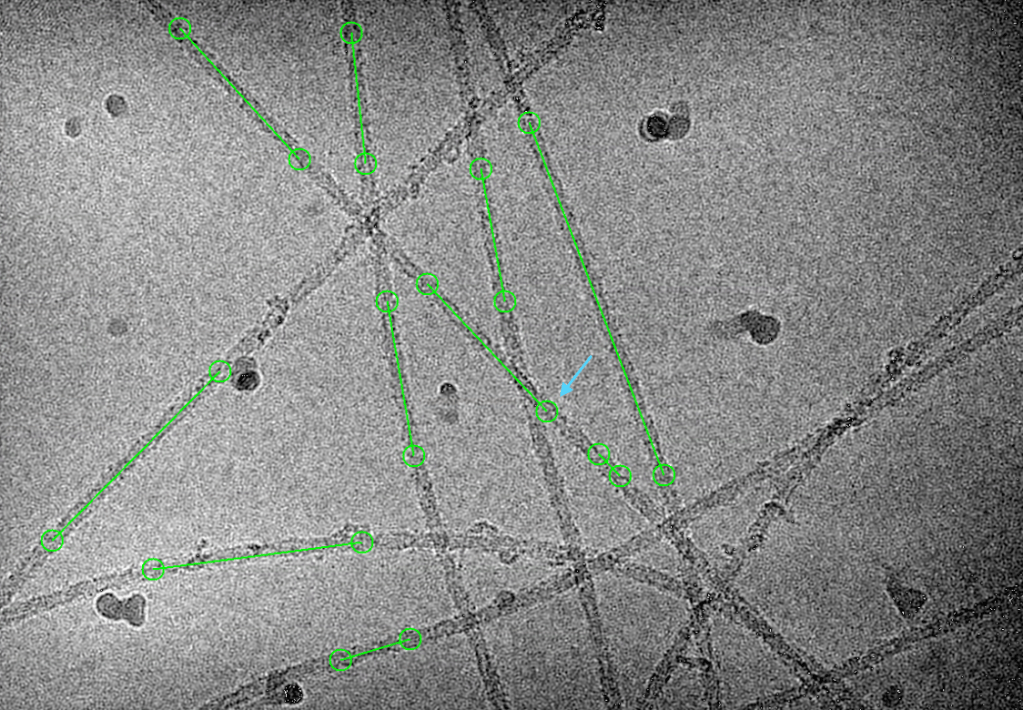
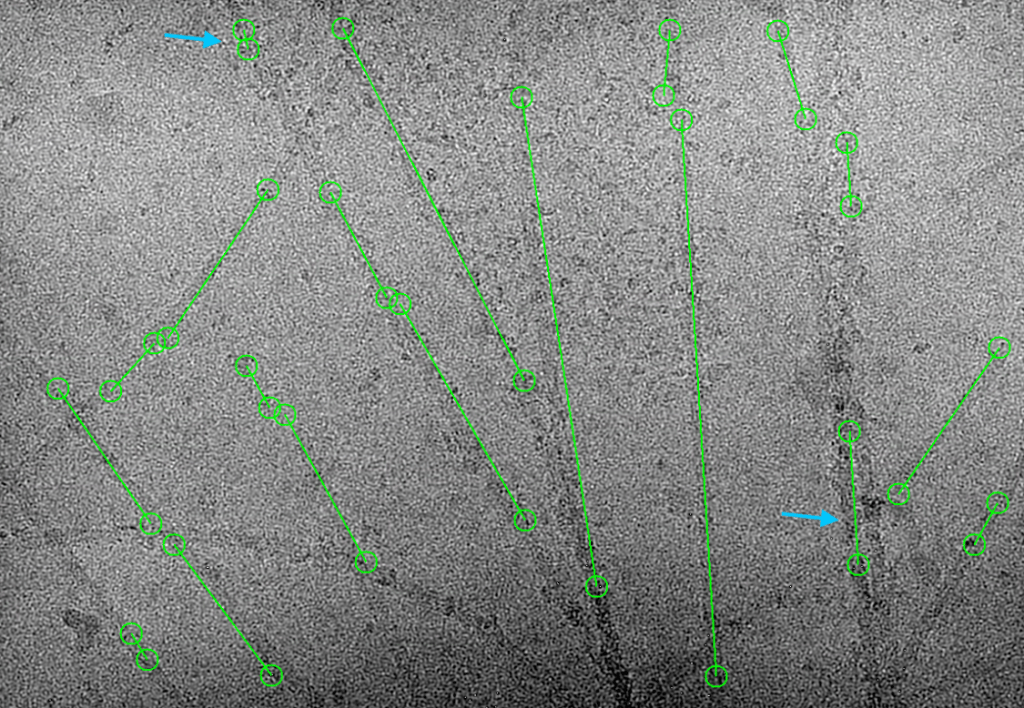
Here are some good and wrong examples crYOLO picked. I think overall goodness is 85%. Considering 5 hours of auto-picking compared to 10-day human picking, the picked fibrils are good enough to move forward next steps.