“alter” is a useful function in PyMOL. One can use “alter” to renumber residues, rename chain IDs, re-define secondary structures et al. More details can be found at PyMOLWiki.
Here I am showing a simple example that using “alter” to redraw secondary structure of ubiquitin (PDB: 1UBQ).
Example commands: (two steps, alter object, then run "rebuild") alter 40-45/, ss='L' rebuild (see second image. the beta strand is shown as loop, cartoon mode) alter 40-45/, ss='H' rebuild (see third image, the beta strand is shown as helix, cartoon mode)
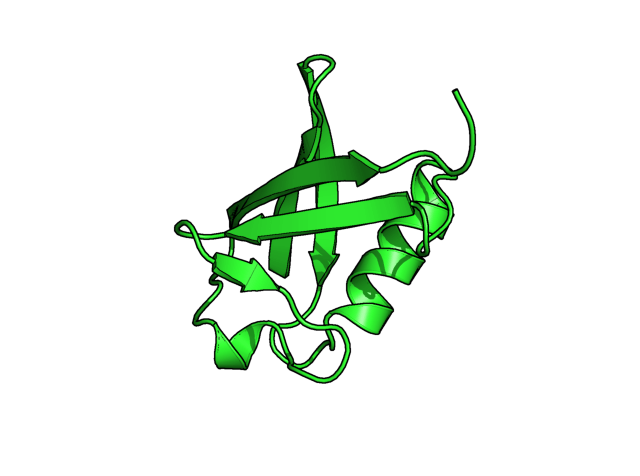

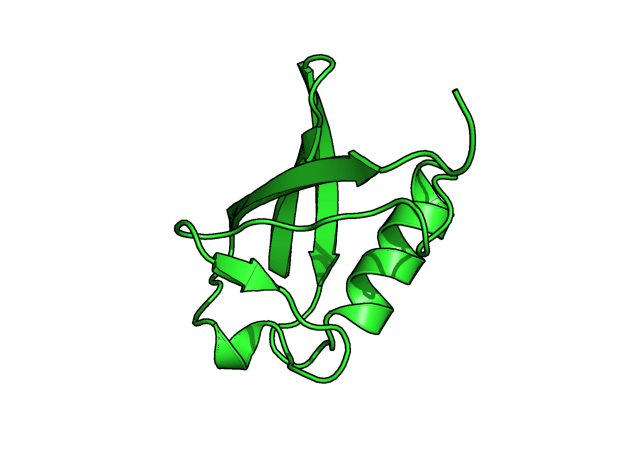

More example commands of alter: (A), alter chainID alter (chain A),chain='B' sort (B), alter residue numbers of chain A alter (chain A),resi=str(int(resi)+1000) sort